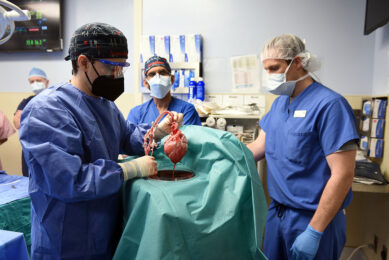
A comparison of individual versus pooled sample testing on pool-level and case-level PRRS virus PCR detection sensitivity and specificity
The time it takes to begin producing porcine reproductive and respiratory syndrome virus (PRRSv)-negative groups of weaned pigs following the new introduction of a non-resident PRRSv and related onset of a clinical reproductive PRRS outbreak in swine breeding herds has become an important means by which to assess the effectiveness of PRRS interventions applied to breeding herds.
The measure “time-to-negative-pigs” (TTNP) is typically described as the number of weeks between the start of an intervention protocol (e.g., herd closure to new replacement introductions and/or mass vaccination) and the first “consistent” production of PCR-tested negative groups of weaned pigs.
For estimating TTNP, the key measurement components are: sampling frame (e.g., piglets within 1 week or less of weaning), sample selection (e.g., smallest barrow from gilt litters, one piglet per litter) sampling method (jugular venipuncture, ear notch), sample type (e.g., blood-to-serum, swab-to-saline), sample size, sampling interval, sample pooling (e.g., pooling of N individual samples), shipping materials and method, PCR assay, laboratory procedures, and the number of consecutive testednegative samplings.
The objective of this study was to assess the effect of sample pooling of 5:1 and 10:1 on PCR detection sensitivity and specificity at both the pool and case levels when estimating TTNP.
Dale Polson1 Greg Hartsook1 Brent Carmichael2
1. Boehringer Ingelheim Vetmedica, Inc., St Joseph, MO, USA;
2. Iowa State University, Ames, IA, USA
For full presentation see attached pdf: