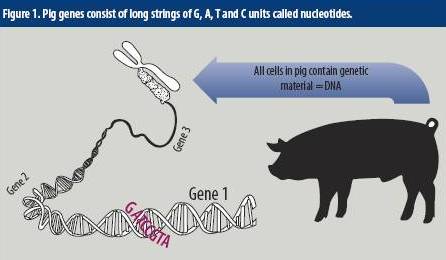
Genotyping: How useful is it for producers?
In the dairy cattle industry, terminology like SNPs, genomic selection and genomic breeding values have already been around for some years now. In the swine industry, technology related to sequencing of the pig genome has started somewhat later but is now well underway. Where are we and what prospects does the technology offer for the industry?
By Christopher K. Tuggle and Jack C.M. Dekkers, Department of Animal Science, Iowa State University, USA
The sequencing of the pig genome will be completed late in 2009, although the analysis of the data is on-going and a complete understanding of what all these data mean is many years off. However, pig producers can already start to make use of the current sequencing data, and what is important to understand now is the value of the genome sequence in developing new genotyping tools to track variation in genes and gene regions that contribute to traits of interest.
First, what do we mean exactly by ‘sequencing the genome’? A genome is made up of specific units of genetic information called ‘nucleotides’ which have four flavours; A, T, C and G. Each gene on a chromosome in a pig cell consists of thousands of these nucleotides, and the identity and function of each gene is due to the specific sequence of these nucleotides along the chromosome (Figure 1).
As genes are discrete locations along a chromosome with spaces of DNA between genes, it is important to sequence the whole chromosome so that we can identify and map where all the genes are. Genome sequencing then consists of determining the exact sequences for all genes and the spaces between genes across each chromosome. The pig whose genome is being sequenced is a single Duroc sow, so we will have only the complete data for her genome. However, this information is very valuable, as it provides a detailed roadmap for where all genes are located, as well as an example sequence for each gene.
Once we have the exact sequence for each pig gene, we can compare those sequences to the human genome sequence to leverage all the biological knowledge regarding the actions of genes. Such ‘comparative’ information is very helpful in selecting genes to study in the pig, as we expect that genes in the pig with very similar sequences to a specific human gene will share the function in disease, growth, or reproduction that has been determined for other species.
Value to breeders
In addition to the one sow being completely sequenced, researchers have also partially sequenced many other animals representing a number of breeds in several countries. This was done to find where in the genome there are differences between breeds and between animals; such differences are called genetic variants. Most genetic variation between individuals is due to differences at a single nucleotide in a stretch of nucleotides, and this
variation is called a ‘single nucleotide polymorphism’ or ‘SNP’ (Figure 2).
variation is called a ‘single nucleotide polymorphism’ or ‘SNP’ (Figure 2).
Researchers have compared such partial genome sequences and have so-far found several hundred thousand SNPs in the genome, both within genes and outside of genes, where the sequence in one animal differs from other animals. Some of this variation is critically important, for example the genetic variant that causes swine stress syndrome is a SNP in the ryanodine receptor gene (RYR1) that affects the function of the protein. Other SNPs may not create a mutation in a gene but can serve as a marker for a specific region of a chromosome that contains genes that are important for specific traits.
So we have a genome sequence of one animal and bits of sequences from many other animals that show SNPs from animal to animal. What can we do with that? With this information, it is possible to determine the exact sequence of an individual pig at a specific location in the genome. This process is called genotyping, as we are determining the specific type of the gene sequence at that position in the genome.
Importantly, each cell in an animal contains two copies of each chromosome, one that the animal received from its sire and one that it received from the dam. The exception to this rule is the sex chromosomes, for which males have only a single copy of the X and Y chromosomes. The ‘genotype’ of an animal at a particular SNP will thus consist of two letters (A, T, G, or C), representing the sequence at that SNP position, one for the chromosome the pig inherited from its sire and one for the chromosome it inherited from its dam. Using the classic Swine Stress gene again as an example, the important variant that causes halothane sensitivity is a SNP at position 1843 in the RYR1 gene, and the two possible nucleotides that can be present at that position in the sequence are T or C. Within a pig, the paternal copy of the RYR1 gene can be the same as or different from the maternal copy. Thus a pig can be genotyped as TT (normal), TC (a carrier) or CC (an affected pig) for the RYR1 position 1843 SNP.
Entire genome
However, genotyping for the stress gene is pretty old history which did not require a pig genome sequence, so why all the fuss? Well, the new information covers the entire genome of the pig, thus we can genotype – and keep track of – many more places in the genome. In fact, a recent tool has been developed that can determine the genotype of a pig at 60,000 different positions across the genome in one shot. This tool is called a SNPChip, and has been in use since late 2008 for genotyping the pig. In most cases, these SNPs are not known to directly contribute to differences in pig traits. Instead, these SNPs were chosen to ‘mark’ a region of a chromosome so that researchers can follow that genome region from animal to animal.
This is valuable because if specific traits of interest have also been recorded on these animals, geneticists can match up the animals with best trait values with specific regions of the genome which those superior animals carry using these ‘marker SNPs’. These SNPs were chosen both to evenly cover the entire genome and based on the ‘informativeness’ of the SNP. This informativeness is based on extensive sequencing data, which indicated that this particular SNP variation could be found in many pig breeds. Thus typing this SNP would likely be ‘informative’ in most population studies.
However, not all SNPs are useful in every population, as some variants are only found exclusively in one breed, or in a small number of breeds. As well, a given variant (e.g. T for the stress gene) may be much more prevalent in one breed as compared to others. However, the large number of SNPs on the SNPChip means that in most populations, the SNPChip will provide enough useful SNPs for many applications.
There are several uses for broad-scale genotyping using the SNPChip. One use is parentage testing, and 60,000 genetic tests across the genome would clearly provide sufficient information to both exclude parentage and provide a measure of certainty in cases of matching genetics. As indicated above, another use for SNPChip data is in mapping the location of genes controlling pig traits. With SNPs marking every region of the genome at a high density, a statistical association of specific regions with favourable traits is possible. This can be done by simply comparing the average trait phenotype of pigs that have, e.g., genotype AA at a specific SNP to the average phenotype of pigs that have genotype AG or GG. When specific regions are thus identified, they can be selected for in populations or introgressed (genetically integrated) into other populations by using only the SNPmarker information. This approach to developing genetic tests for specific traits has been used before and has led to, e.g. the Estrogen Receptor Gene test for litter size and the MC4R test for feed intake, growth and efficiency, and several of such individual tests are commercially available. Although these genetic tests are useful, they typically explain only a limited part of the genetic differences between individuals. Thus, although they can help identify the better animals, regular estimates of breeding values based on phenotype remain essential to making good selection decisions. However, with the new SNPChip, we can now conduct these tests at many more locations in the genome.
This has the potential to have genetic tests across the genome that, together, explain a much larger part of the genetic differences between individuals. And perhaps give sufficient accuracy that selection can be based on this combination of genetic tests alone. This process of using many genetic tests across the genome in selection is what is called “genomic selection” and will be explained in further detail below.
Using genetic tests
What is whole genomic selection and how can it be used in the industry? The main principle of genomic selection is outlined in Figure 3. The first step in genomic selection is to collect phenotypes and DNA from a large group of pigs from a population and to genotype each animal for all SNPs on the SNPChip. The resulting ‘training data’ is used to ‘train’ a statistical model that estimates the effect of each of the SNPs on the SNPChip with the trait phenotype.
In principle, the estimate for a given SNP is based on the comparison of average phenotypes of individuals that have alternate genotypes at that SNP, as described above, but in genomic selection this is done simultaneously for all SNPs on the SNPChip. The resulting estimates can then be used to predict the ‘genomic’ breeding value of new individuals based on their genotypes for the SNPChip, as illustrated in Table 1 for a simple example with three SNPs. Expectations are that, with a large enough training data set (several thousands of individuals), and depending on heritability of the trait, genomic selection can lead to estimates of breeding values that are more accurate than those that would be obtained from observing the pig’s own phenotype or that of its relatives. Thus, genomic selection could in principle allow selection of pigs at a very young age, without having to wait to record their phenotypes.
Status and prospect
Estimates of breeding values based on genomic selection have become available to the dairy cattle breeding industry in the past year in several countries. The main use of the genomic breeding values has been to genotype young bull calves to decide which should be entered into progeny testing programmes. Some AI bull studs have also used genomic breeding values to market semen from young bulls prior to them having completed their progeny test evaluation. Genotyping of heifers and cows to determine those that should be used for breeding is also gaining momentum. The prospect for genomic selection in dairy cattle is that it will lead to much shorter generation intervals because accurate estimates of breeding values can be obtained at a much younger age. This is expected to lead to substantial increases in the rate of genetic improvement.
The SNPChip for pigs has not been available very long, so the implementation of genomic selection is not yet at the same stage as it is in dairy cattle. In addition, the current genotyping cost of around US$250 per animal will likely be more prohibitive in pigs than it is in dairy cattle, because of the greater value of the individual breeding animal in cattle. Nevertheless, there are important opportunities for genomic selection in pigs also. Most of these opportunities lie within the elite breeding herds and populations that create genetic progress, rather than in individual producer herds.
Benefits will be limited for selection for growth rate and backfat, because these traits can be recorded relatively early in a pig’s life and without tremendous costs. However, genomic selection would be useful in particular for traits that can only be recorded late in life or only directly on one sex (e.g. litter size); for meat quality traits that require traits to be recorded on sibs or progeny that are slaughtered; or on disease resistance traits. In addition, genomic selection may allow more effective selection for performance in the field, instead of for performance in a high-health nucleus herd environment by ‘training’ the prediction model on genotypes and phenotypes that are collected in the field.
In conclusion, although genomic selection is still at its infancy in pigs, opportunities exist for enhancing genetic improvement programs in pigs, in particular for traits in which selection is currently difficult. In any case, the application of the new pig genome sequence data in high density SNPChip genotyping of pigs is certain to expand in the future.
Source: Pig Progress Volume 25 nr 9











