Sequenced pig genome – past and future tales

Last year the first pig genome was sequenced by an ?international consortium. Having a reference, it is easier to add 48 others. The genetic diversity provides the pig industry with ?interesting stories of the pig’s history – and shows paths for future research.
By Prof Martien A.M. Groenen, Animal Breeding and Genomics Centre, Wageningen University, the Netherlands
Genetic variation within a species or population is the key to animal breeding as it provides the raw material on which future progress in breeding depends. In pigs, for centuries man has used this variation to develop an animal targeted to his needs and wishes. This process goes back as far as 10,000 years ago, the estimated start of the domestication process of the pig. For most of this period selection was empirical and directed by, what Darwin refers to as, unconscious selection. It was not until the late 18th and 19th century and in particular after the 1950s, that pig breeding increasingly became based on scientific genetic theory.
Today modern pig breeding is a technology-based industry that makes use of the latest advancements in computing technology, genetics and genomics. The increasing interest in genomics in pig breeding goes back to the late 1980s and early 1990s due to developments in molecular biology and in particular the start of the human genome sequencing project. Funded by the European Union (EU) and the United States Department of Agriculture (USDA), in both Europe and the US researchers initiated systematic mapping of the porcine genome resulting in detailed genetic maps in the mid-1990s. Then, in September 2003, academic, government and industry representatives convening in Jouy-en-Josas near Paris formed the Swine Genome Sequencing Consortium (SGSC) to provide international coordination for sequencing the pig genome. Finally in November 2012 the SGSC, coordinated by Larry Schook (University of Illinois, USA), Alan Archibald (Roslin Institute, UK) and myself, reached its major goal by publishing the first draft genome of the pig in the leading scientific journal Nature accompanied by over 20 other publications in other journals covering further aspects of the genome of this interesting animal.
Genetic maps
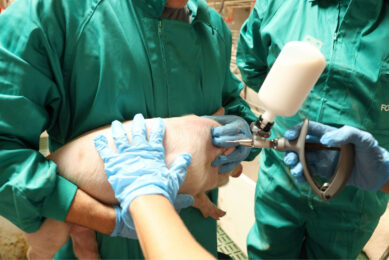
Pig geneticists have come a long way in these past 20 years since the earliest efforts to generate genetic maps of the porcine genome. At that time nobody could have even imagined where we would be today. While describing the genome of a single Duroc animal as the ‘golden’ reference for the pig’s genome, the Nature paper also presents the analysis of the completely sequenced genomes of another 48 pigs from a variety of breeds and wild boar populations in Europe and Asia. Once a reference genome is available, this acts as the golden standard, or reference, to which all the other genomes of the same species are compared. Consequently, the costs to sequence any additional individual are several orders of magnitude lower than was required for that first individual. For pigs, this first individual was T.J. Tabasco (see picture) a female domestic Duroc pig born in 2001 in Illinois, USA. The choice for her is easy to explain – much material had already been stored, as that had been sequenced in earlier projects making clones.
New sequencing technologies have resulted in an ongoing increase in the rate at which individual genomes are being sequenced with a continued decrease in sequencing cost (Figure 1). In a study financed by the European Research Council, the group at Wageningen University currently has already sequenced close to 250 individual pigs, and the total number of pigs sequenced worldwide is most likely approaching 500. So, what have we learned from these genomes and how is this going to affect pig breeding and the use of the pig in other research areas in the future?
Genome evolution
The pig genome is of similar size, complexity and organisation as the human genome. It has around 22,000 genes a number similar to that of human and other mammals and in the pig these genes are located on 19 different chromosomes. Comparing these genes of the pig to those of other mammals like the dog, horse, mouse and human, reveals that some gene families are undergoing relatively fast evolution in the pig, with particular types of genes rapidly expanding. Examples are immune genes and, perhaps not surprisingly, olfactory genes. The pig has more unique olfactory genes than humans, mice or dogs. In fact, the pig has the largest number of active olfactory genes seen in any mammal whose genome has been sequenced thus far, showing the importance of smell in this scavenging animal. And while pigs can smell a world of things humans and many other animals can’t (like e.g. truffles), their sense of taste is somewhat impaired. Pigs have significantly fewer bitter taste receptor genes than humans, for example, and genes involved in perception of sweet and umami flavours (which humans perceive as meaty) are also different in pigs and humans.
Pigs come in a wide variety of breeds and anyone who has seen Asian pig breeds is aware of the very different morphology of these pigs compared to European breeds. But how different are these pigs actually? While many of these differences probably are due to human intervention by breeding for different characteristics in Europe and Asia, over the past there has been growing evidence for the long divergence between European and Asian pigs and that these breeds originated from wild boars through independent domestication events in Europe and Asia (further discussed below). The comparison between the genomes of six European and four Chinese wild boars revealed significant genetic differences, and it was estimated that European and Asian wild boar separated from one another roughly one million years ago.
Other information that could be extracted from the genomes of these wild boars was that both the European and Asian wild boar lost much genetic diversity some 20,000 to 50,000 years ago, likely as a result of global glaciation events. The last ice age, which reached its maximum around 20,000 years ago, resulted in a strong reduction and fragmentation of the habitat of the pig across Eurasia but this effect was most severe in Europe. The result was a strong reduction in effective population size of the pig and this effect was much larger in Europe than in Asia. These effects are clearly visible in the genomes of these wild pigs even today, with lower genetic variation in the European wild boars compared to its cousins living in Asia.
Domestication and selection
The genome sequence of the European and Asian wild boars confirmed earlier findings based on studies that focused on the mitochondrial DNA, a small DNA molecule only inherited from the mother, that pigs in Europe and Asia were domesticated completely independently of each other. This probably already resulted in very distinct different types of domesticated pigs in Europe and Asia. While in Europe until the late middle ages pannages were used where pigs were freely roaming the forest and fed on mast, in Asia pigs were held in enclosures and fed on scraps and human refuge. Consequently, domesticated pigs in Europe most likely still had many of the characteristic features of wild pigs, while in Asia pigs were characterised by their round potbellies and short legs. European pig farming changed during the industrial revolution and Asian domesticated pigs were used to improve the European breeds in the 18th and 19th centuries, a process well documented. The analysis of the genome of individual pigs from a variety of commercial breeds of European origin revealed that a surprisingly large fraction of their genome could be traced to these introgression events in the 18th and 19th century. Based on the genome sequence of individual Duroc, Large White, Landrace, Pietrain and Hampshire pigs an estimated 35% of the genome of today’s commercially bred pigs can be traced back to these original Asian breeds.
Phenotypic changes
All these events have resulted in a large number of drastic phenotypic changes in domestic pigs compared to its ancestor the wild boar for traits like body composition, morphology, coat colour and reproduction. By comparing the genomes of domestic pigs with that of wild boars my group in collaboration with Leif Andersson (Uppsala University), Merete Fredholm (University of Copenhagen) and Alan Archibald, identified three loci that affect one of the most striking morphological changes in the domestic pig – the elongation of the back and the increased number of vertebrae. This striking change had already been observed and described by Charles Darwin in his in 1868 published book The variation of animals and plants under domestication. We were able to identify three genes located at these loci – NR6A1, PLAG1 and LCORL – that are strong candidates for the observed changes in this trait in the European domestic pig. Interestingly two of these genes – LCORL and PLAG1 – have also been associated with variation in size in other species including humans.
The analysis of selection for different coat colour phenotypes in pigs provides an excellent example of the complex nature of some of the changes in the genome that underlie phenotypic differences. The variants at the KIT locus, long known to affect coat colour phenotypes in the pig including dominant white, patched and belted, were shown to be the result of a complex organisation of multiple independent duplication events (Figure 2). These findings suggest that haplotype effects due to multiple sequence polymorphisms affecting the structure or expression of a single gene may be common in many species and are probably one reason why it is so difficult to reveal the causative mutations at loci underlying complex traits and disorders. Such complex structural changes that affect a phenotype may be much more common than so far realised, something that also is highly relevant for studies on complex diseases in humans.
The future
Over the centuries pigs have been an important source of animal protein. Production and consumption amounts to a 100 billion kg per year, and makes pork the most widely eaten meat in the world today. Pig production and pig breeding is constantly facing new challenges due to changing consumer demands, environmental influences including specific diseases and changes required because of increased emphasis on animal welfare and animal health.
Already by the end of 2008, the then ongoing sequencing of the porcine genome enabled the development of a porcine SNP chip with 60,000 genetic variants (single nucleotide polymorphisms; SNPs) that has and still is being used extensively for genomic selection by the pig breeding industry. The sequencing of the genome of different pigs has shown that the variation in the porcine genome, even within most commercial breeds, is two to three times higher than in humans. The domestic pig still has an ancestor-like wild cousin on the loose, unlike the domestic cow e.g., whose ancestors, the aurochs, are now extinct. The porcine lineage has a lot of genetic diversity remaining. In the hundreds of individuals sequenced to date we have already identified over 30 million genetic variants, clearly showing the available potential for further improvements of these breeds in the future.
The publication of the pig genome in 2012 clearly reflects a major milestone in pig genetics. Nevertheless it only marks the beginning of a highly, much more detailed map and understanding of the pig genome, in which hundreds, if not thousands, of animals will play a part. This information will be invaluable not only to the pig-breeding sector, but also in terms of using pigs for biomedical and fundamental biological research. We already found many variants of genes that may be linked to an increased risk of certain diseases in humans, such as obesity, diabetes, Alzheimer’s and Parkinson’s disease. We can now start studying the effect of these variants in animals that are physiologically similar to humans. Like humans, pigs are omnivorous and their digestion and physiology are very similar to that of humans. In addition, the size of their internal organs, including the heart, kidneys and pancreas, largely corresponds with the size of human organs. This makes the pig an important model for medical research as well.