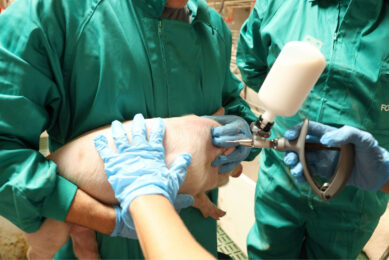
MRSA not influenced by pigs’ nasal microbiota

Tests with liquid and conventionally fed pigs show that no influence has been found of nasal microbiota on MRSA.
This was reported by researchers at the University of Guelph’s Ontario Veterinary College, Canada. Farm management, however, can make a difference, they said.
In a research published in BMC Veterinary Research, the scientists write that nasal microbiota, although poorly assessed, could play a role in carriage of important micro-organisms such as methicillin-resistant Staphylococcus aureus (MRSA).
The objectives of their study were to describe the nasal microbiota in slaughter age pigs, to evaluate the impact of farm management on the nasal microbiota and to provide a preliminary assessment of the influence of the microbiota on MRSA carriage.
Results
Nasal swabs were collected from five MRSA positive and eight MRSA negative pigs on one farm that used a liquid feeding system and routine tylosin treatment, and seven MRSA negative pigs from an antibiotic-free farm that used conventional feeding.
A total of 946,310 sequences passed all quality control filters. The number of sequences per sample ranged from 4,307 to 165,656.
Proteobacteria predominated in all but two samples. Liquid-fed/ tylosin-exposed pigs had significantly lower relative abundances of Verrucomicrobia, Fibrobacteres and sequences unclassified at the phylum level.
When comparing only liquid-fed pigs, MRSA carriers had significantly more Bacteroidetes than MRSA negative pigs.
In total, 124 genera were identified, with Moraxella accounting for 35.4% of sequences. In the Jaccard index tree, five of eight MRSA positive pigs clustered closely together, as did six of the seven conventionally-fed pigs.
A significant difference was identified between conventional and liquid-fed pigs using parsimony test with the Jaccard but not the Yue & Clayton index. There were no significant differences between MRSA positive and negative pigs.
Operational taxonomic units (OTUs) belonging to Firmicutes were the main indicators of MRSA negative pigs, including Lactobacillus and another Lactobacillaceae and Staphylococcus.